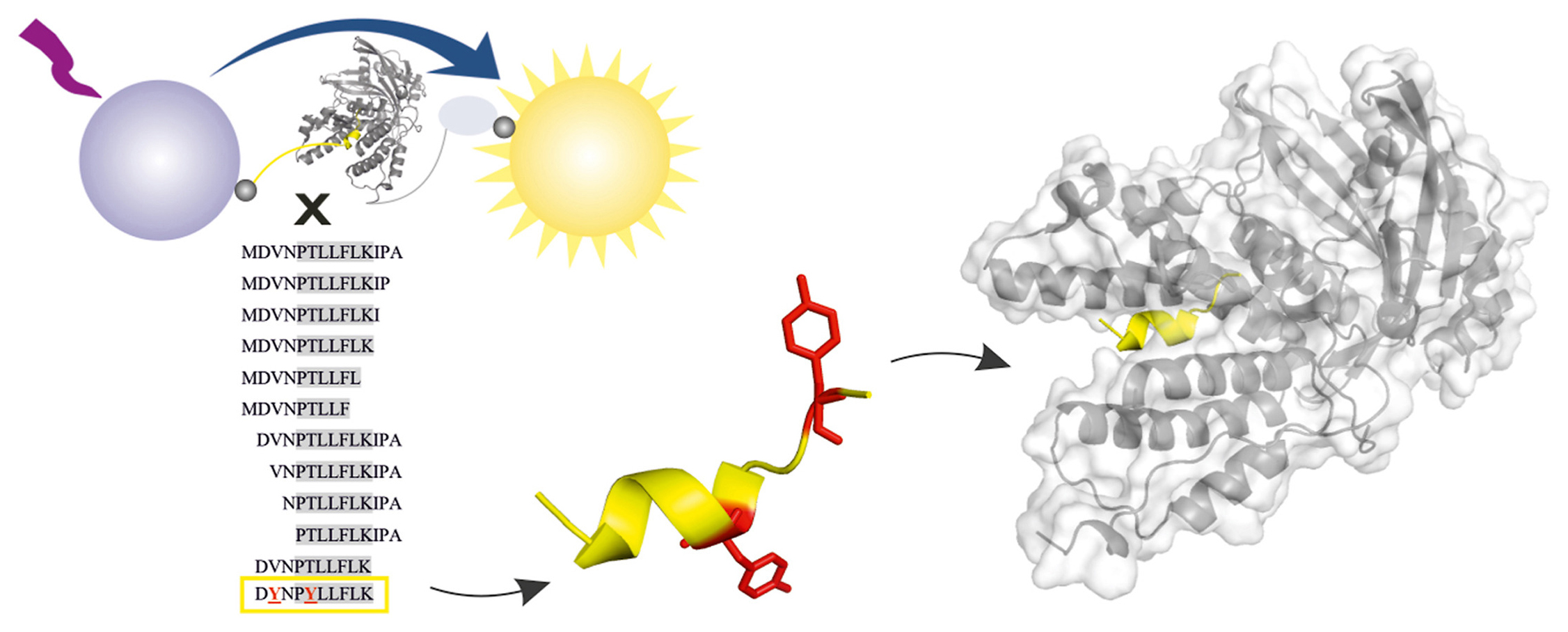
Structural characterization of the interaction between the C-terminal domain of the influenza polymerase PA subunit and an optimized small peptide inhibitor
We have a new paper in Antiviral Research. Congrats to Milan, Jakub, and the whole team!
Citation: Hejdánek, J.; Radilová, K.; Pachl, P.; Hodek, J.; Machara, A.; Weber, J.; Řezáčová, P.; Konvalinka, J.; Kožíšek, M. Structural characterization of the interaction between the C-terminal domain of the influenza polymerase PA subunit and an optimized small peptide inhibitor. Antiviral Research 2021, 185, 104971. https://doi.org/10.1016/j.antiviral.2020.104971
Abstract: Influenza viruses can cause severe respiratory infections in humans, leading to nearly half a million deaths worldwide each year. Improved antiviral drugs are needed to address the threat of development of novel pandemic strains. Current therapeutic interventions target three key proteins in the viral life cycle: neuraminidase, the M2 channel and RNA-dependent-RNA polymerase. Protein-protein interactions between influenza polymerase subunits are potential new targets for drug development. Using a newly developed assay based on AlphaScreen technology, we screened a peptide panel for protein-protein interaction inhibitors to identify a minimal PB1 subunit-derived peptide that retains high inhibition potential and can be further modified. Here, we present an X-ray structure of the resulting decapeptide bound to the C-terminal domain of PA polymerase subunit from pandemic isolate A/California/07/2009 H1N1 at 1.6 Å resolution and discuss its implications for the design of specific, potent influenza polymerase inhibitors.